Latest Research
- 2022.03.01
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
- Ueda-Kitaguchi Group
Handy fluorescence labeling of antibody proteins with coiled-coils
―High performance sensor antibodies can be selected using baker's yeast―
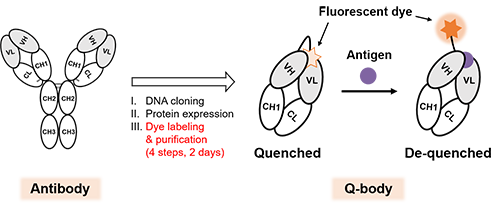
Immunoassays that can allow us to specifically detect trace substances in samples taking advantage of antibodies are widely used in various fields such as clinical diagnosis, food hygiene, and biological research. Although high-molecular-weight antigens such as proteins and viruses are detected with high sensitivity, it is difficult to measure low-molecular-weight antigens with high sensitivity. On the other hand, Quenchbody (Q-body) (Fig. 1)[Ref. 1], which we have previously constructed by fluorescently labeling the N-terminal region of the antibody, enables measuring the amount of antigen with high sensitivity and ease for both high- and low-molecular-weight targets. Q-bodies are based on the principle of fluorescence augmentation upon antigen binding while the fluorescent dye is quenched in the absence of antigen. Because of its ease of handling, Q-bodies are expected to be applied as a core technology for rapid and easy diagnostic kits. However, the current Q-body construction method takes time and effort. Hence a method that can rapidly convert antibody fragments into Q-bodies is awaited to efficiently conduct studies necessary to improve the antigen response of the Q-bodies.
 |
|
| Fig. 1. | Schematic of antibody and Q-body. The Q-body increases its fluorescence intensity upon antigen binding, enabling high-sensitive detection of both high- and low-molecular-weight antigens. However, the construction of Q-body from recombinant antibody fragments is time-consuming (4 steps, 2 days). |
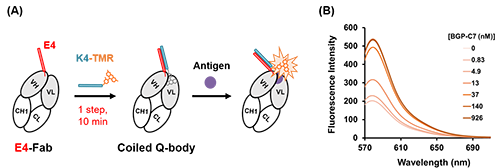
In this study, we focused on the coiled-coil forming peptide pair E4 and K4, which forms a strong heterodimer, and succeeded in rapidly converting antibody fragments into Q-bodies[Ref. 2]. In other words, by mixing an anti-BGP (bone Gla protein, a bone disease marker) Fab fragment with E4 fused to the N-terminus of the heavy chain and K4 labeled with the fluorescent dye TAMRA, a Coiled Q-body (CQ-body) (Fig. 2) could be constructed from antibody fragment in one step and within 10 minutes without purification.
 |
|
| Fig. 2. | (A) Schematic of CQ-body construction using the coiled-coil forming peptide E4 and K4. The CQ-body could be obtained without purification by just mixing E4-Fab and K4-TAMRA within 10 minutes. (B) Fluorescence spectrum of CQ-body with antigen BGP-C7 (C-terminal 7 amino acids peptide of BGP). |
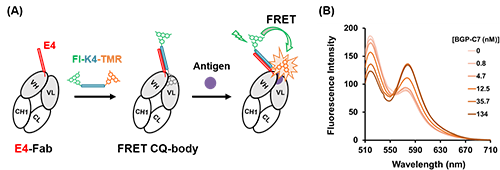
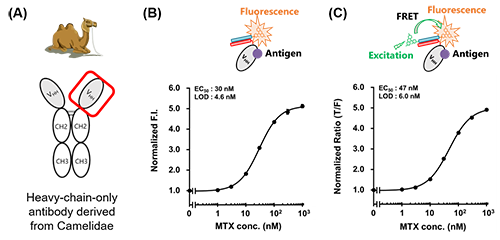
We also succeeded in constructing FRET type CQ-body that enables more precisely antigen detection by a ratiometric assay based on FRET (Förster resonance energy transfer) by introducing FITC as a fluorescent donor into the K4-TAMRA (Fig. 3). Interestingly, a decrease in donor fluorescence and an increase in acceptor fluorescence were observed, which would be caused by antigen-dependent moving out of the TAMRA from the antibody, suggesting the possibility of enabling antigen measurement by changes in fluorescence color using FRET CQ-bodies. Furthermore, these CQ-bodies or FRET CQ-bodies were successfully constructed not only with the anti-BGP Fab fragment but also with anti-MTX (methotrexate, an anticancer drug) VHH, a heavy-chain-only antibody fragment derived from Camelidae, with a response comparable to that of Q-bodies constructed by conventional methods (Fig. 4).
 |
|
| Fig. 3. | (A) Schematic of FRET CQ-body construction using dual-color K4 peptide. (B) Fluorescence spectrum of FRET CQ-body with antigen BGP-C7. |
 |
|
| Fig. 4. | Schematic of Camelidae-derived heavy-chain-only antibody (A) and detection of MTX by VHH-based CQ-body (B) or FRET CQ-body (C). |
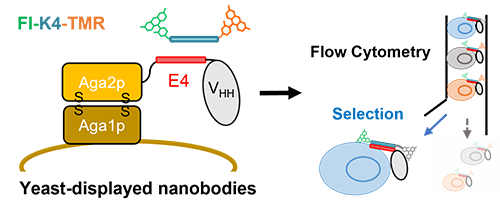
Next, we aimed to construct the FRET CQ-body on yeast cells by using yeast surface display [Ref. 3] to select the potent nanobody-based Q-bodies (Fig. 5). Initially, yeast-displayed anti-MTX VHH was successfully converted to a Q-body and showed significant quenching and MTX-dependent dequenching, which was detected by flow cytometry (Fig. 6). Also, we succeeded to select highly responsive nanobody-based Q-bodies from a small nanobody library consisting of 30 variants that recognize HSA (human serum albumin), which demonstrates the applicability of the developed method (Fig. 7). However, the fluorescence response of selected nanobodies was relatively modest when expressed by E. coli. compared with that on the yeast cell surface.
 |
|
| Fig. 5. | Schematic image of yeast-displayed nanobody-based Q-body and its selection steps |
 |
|
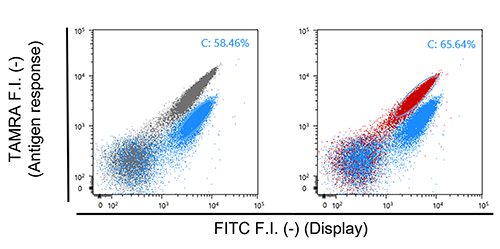
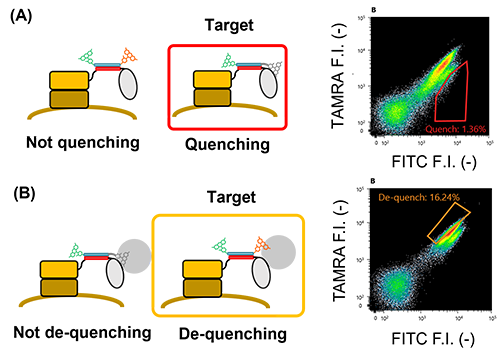
| Fig. 6. | Flow cytometric analysis for yeast cells displaying E4-MTX VHH labeled with FITC-K4-TAMRA. (A) Analysis of quenching. (B) Analysis of de-quenching. Grey dots represent the yeast cell displaying E4 peptide. Blue dots represent the yeast cell displaying MTX VHH in the absence of MTX. Red dots represent the yeast cell displaying MTX VHH in the presence of MTX. |
 |
|
| Fig. 7. | Selection of potent anti-HSA nanobody-based Q-body displayed on the yeast cell surface for quenching clones (A) and de-quenching clones (B). Quenching clones were selected by the red gate and de-quenching clones were selected by the yellow gate. |
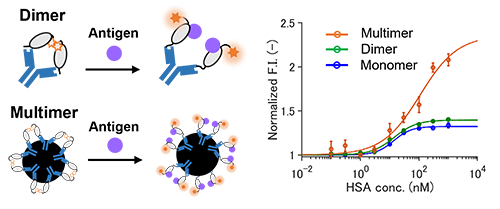
Therefore, we aimed to mimic the yeast surface display by using beads for protein purification or IgG antibody (Fig. 8). Interestingly, when the Q-body was multimerized by beads, it showed a higher response than monomeric Q-body, which indicates that we have indeed succeeded to select a potent nanobody-based Q-body; sometimes the response could be improved by multimerization [Ref. 4].
 |
|
| Fig. 8. | Detection of HSA by Q-body using selected nanobody at the state of one molecule (monomer), two proximate molecules (dimer), multiple molecules captured on beads (multimer). |
These results suggest that this technology will be applied to quickly obtain well-behaved Q-bodies and other fluorescent biosensors for various targets through directed evolutionary approaches.
References
| [1] | R. Abe, H. Ohashi, I. Iijima, M. Ihara, H. Takagi, T. Hohsaka, H. Ueda. J. Am. Chem. Soc. 133, 17386-17394 (2011) |
| [2] | T. Yasuda, A. Inoue, T. Kitaguchi and H. Ueda. Chem. Commun. 57, 8206-8209 (2021). |
| [3] | T. Border & D. Wittrup, Nat. Biotechnol. 15, 553-557 (1997). |
| [4] | A. Inoue, T. Yasuda, B. Zhu, T. Kitaguchi, A. Murakami, H. Ueda, Sci. Rep. 11, 22590 (2021). |



