Latest Research
- 2023.12.01
- Tanaka-Yoshida Group
Essential roles of the redox regulation system in plant photosynthesis
Plant photosynthesis produces organic compounds using light energy from the sun and thereby supports all of the life on the Earth. Because plants cannot escape from ever-changing light environments, they have to regulate photosynthetic activities flexibly under such conditions. Elucidating regulatory mechanisms of photosynthesis is one of key challenges in the field of plant science.
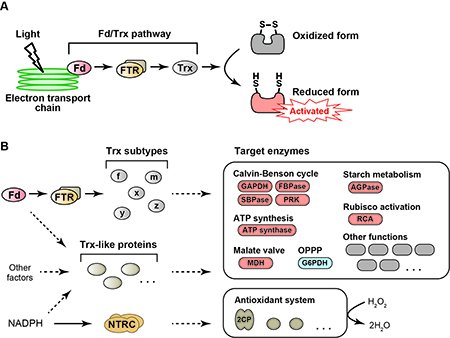
We are focusing on the redox regulation system, a posttranslational protein modification based on the reduction/oxidation switch. This regulatory system is ubiquitously conserved in all organisms from prokaryotes to eukaryotes. Plant chloroplast, a site for photosynthesis, contains this system, but its mode of action is unique in terms of linking to light (Fig. 1A). Upon irradiation, photosynthetic electron transport reactions are triggered in the thylakoid membrane in chloroplasts, generating the reducing power. A part of reducing power is transferred from an electron carrier protein, ferredoxin (Fd), to a key factor of redox regulation, thioredoxin (Trx). Trx then transfers reducing power to its target enzymes in chloroplasts. Target enzymes are basically shifted to the activated state upon the reduction. Some enzymes involved in key reactions of photosynthesis, including ATP synthesis and CO2-fixing metabolism, are subjected to redox regulation. Therefore, the redox regulation system in chloroplasts enables the light-responsive activation of photosynthesis. The redox regulation system based on the Fd/Trx pathway has been classically known as the textbook knowledge (Fig. 1A).
It has been recently suggested that the redox regulation system in chloroplasts is more complex than previously expected (1). For example, five Trx subtypes are present in chloroplasts. Despite the identical active site motif, they have different features in the redox regulation functions. Plant genomic studies also have indicated the existence of putative Trx-like proteins in chloroplasts. Furthermore, advances in the proteomics study have led to the identification of potential target enzymes of redox regulation, which are involved in a wide range of chloroplast functions. The diversity of regulatory factors and target enzymes indicates that the redox regulation system in chloroplasts develops divergent pathways for reducing power and systematically controls chloroplast biology (Fig. 1B). How is the redox regulation system organized in chloroplasts? What is the physiological significance of this system? Our goal is to understand these issues comprehensively. We recently identified the protein-oxidation machinery, which has been unclear for half a century (2,3). We, however, need further studies to elucidate the redox regulation system in more detail.
One of the elusive questions is the biological importance of classic Fd/Trx pathway. We recently addressed this simple but critical issue (4).
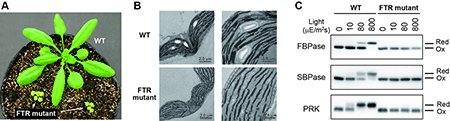
There has been no report of complete suppression of Fd/Trx pathway to date. To overcome this problem, we used the CRISPR/Cas9 gene editing technology. We generated the Arabidopsis mutant plants defective in Fd-Trx reductase (FTR), a central hub in the Fd/Trx pathway. FTR mutant plants showed severe growth impairment, which is accompanied by lowered photosynthetic efficiency and abnormal development of chloroplasts (Fig. 2A, B). We further examined the redox states of several enzymes involved in photosynthetic reactions. In the wild-type plant, these enzymes were shifted to the reduced active state in response to light, while such responses were totally abolished in FTR mutant plants (Fig. 2C). These results clearly indicate that the Fd/Trx pathway is essential for light-dependent activation of photosynthetic enzymes and thus photosynthetic biomass production (4).
This study provides an important insight into the redox regulation system in chloroplasts. We also obtained an unexpected result showing a possibility of the presence of unidentified regulatory pathways. Future studies are required to clarify the overall structure and function of the redox regulation system in chloroplasts.
References
| 1. | Yoshida K, Hisabori T (2023) Plant Cell Physiol 64: 704-715 |
| 2. | Yoshida K, Hara A, Sugiura K, Fukaya Y, Hisabori T (2018) PNAS 115: E8296-E8304 |
| 3. | Yokochi Y, Fukushi Y, Wakabayashi KI, Yoshida K, Hisabori T (2021) PNAS 118: e2114952118 |
| 4. | Yoshida K, Yokochi Y, Tanaka K, Hisabori T (2022) J Biol Chem 298: 102650 |